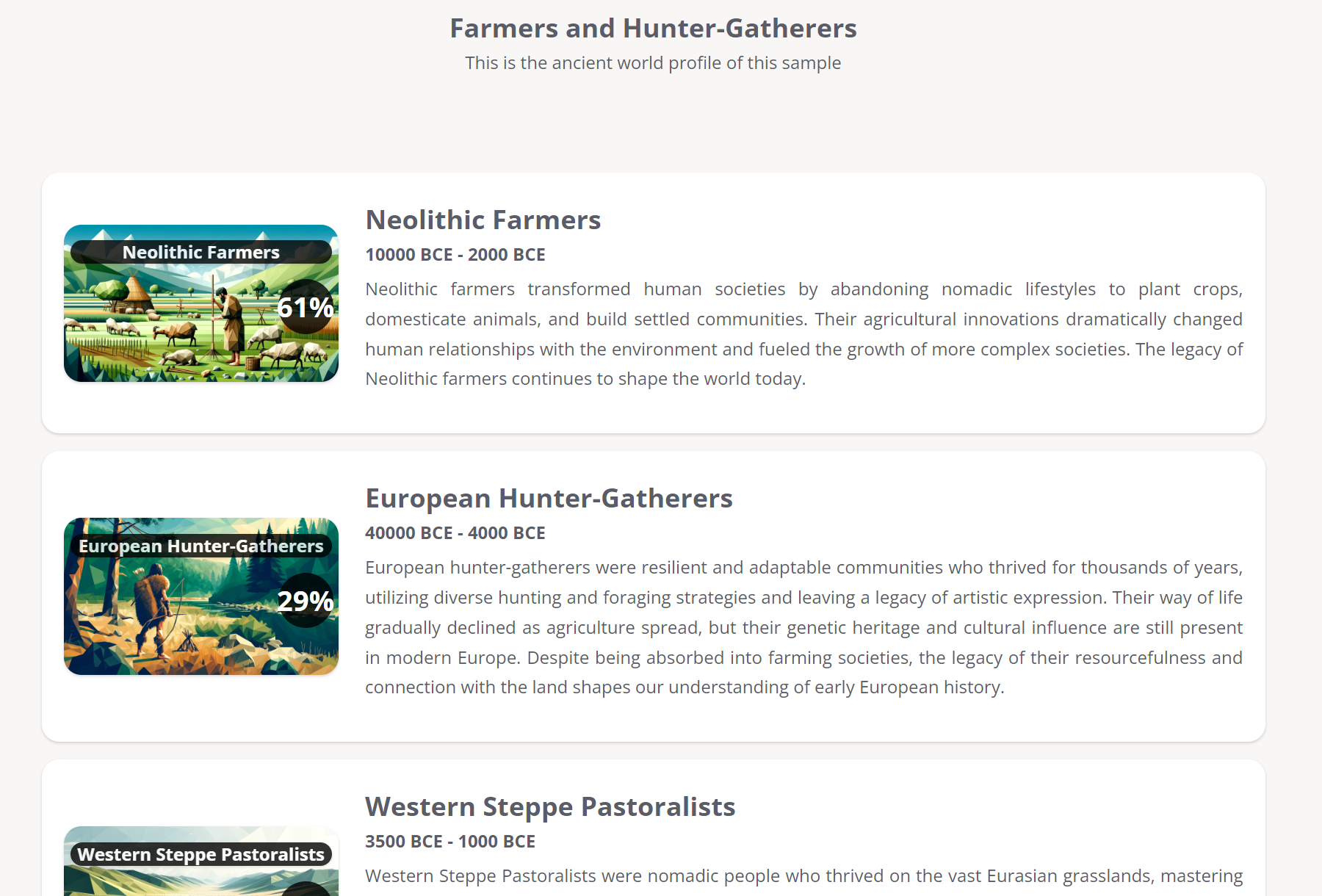
A summary of configurations from our G25 Studio tool.
www.dnagenics.com/g25studio
1. Montecarlo Configuration
This section deals with the core analysis settings:
- Show Sample Name in Header: Displays the sample name in the results header.
- Algorithm: Choose from various algorithms for modeling ancestry:
- Montecarlo V1 and V2
- SLSQP
- BAT (Metaheuristic)
- PSO (Metaheuristic)
- Montecarlo distance: Adjusts the analysis range. Smaller distances include more distant ancestry, larger distances focus on closer ancestry.
- Distance method: Choose between Euclidean, Manhattan, or Chebyshev distance metrics.
- Maximum populations: Limit the number of populations in the analysis (2-10 or all).
- Group populations: Summarizes results for each population group.
- Top 10 montecarlo results: Enables an alternative analysis showing the top 10 Montecarlo results.
- Population proportions: For Top 10 Montecarlo, set custom proportions for result calculation.
2. Oracle Configuration
This section configures the “Oracle” analysis:
- Oracle distance: Similar to Montecarlo distance, adjusts the analysis range.
- Distance method: Includes Angular distance in addition to the Montecarlo options.
- Number of populations: Sets the number of combinations for each Oracle result (10-100).
3. Analysis Tools Configuration
This section enables various data visualization and analysis methods:
- Pearson Correlation: Shows linear relationships between variables. Pearson Correlation is a way to see how similar your genetic makeup is to those of different populations. It calculates a number that shows how closely your genetic data matches with each population group.
If your Pearson Correlation with a specific population is high, it means your genetic makeup is quite similar to that population. If it’s low, it suggests your genetic makeup is less similar to that group.
In mathematical terms, Pearson Correlation quantifies the linear connection or resemblance between two sets of data. It assesses how effectively the data points in these two datasets are interconnected or move in tandem. This measurement spans from -1 (representing a complete negative correlation) to 1 (indicating a complete positive correlation), with a value of 0 denoting the absence of a linear relationship. - K-Means Clusters: Enables cluster analysis with adjustable number of clusters (2-20). K-Means clustering helps us group populations that have similar genetic backgrounds. K-Means can sort these groups into clusters, so you can see which ones are more alike genetically. This can help us learn about their shared history and differences in their genes. It’s like putting puzzle pieces together to understand how people are connected genetically.
- PCA (Principal Component Analysis): Visualizes relationships between samples and populations. Through the use of a PCA chart, we are able to plot your genetics in conjunction with those of various samples in order to understand the degree of genetic proximity between you and the other individuals.
- UMAP (Uniform Manifold Approximation and Projection): Uniform Manifold Approximation and Projection (UMAP) is a non-linear dimensionality reduction technique used in machine learning and data analysis. It’s similar in some ways to other techniques like Principal Component Analysis (PCA) or t-Distributed Stochastic Neighbor Embedding (t-SNE), but it has some unique features and advantages.Key Points about UMAP:
- Preserving Neighborhoods: UMAP focuses on preserving the relationships between data points that are close to each other in the high-dimensional space. This means that if two data points are similar or “neighbors” in the original data, they should still be close to each other in the lower-dimensional representation.
- Non-Linearity: Unlike PCA, which primarily focuses on linear relationships in the data, UMAP can capture non-linear relationships. This is important when your data has complex patterns that can’t be adequately described by linear transformations.
- MDS (MultiDimensional Scaling): Projects pairwise distances between samples. An MDS (Multidimensional Scaling) chart is a visual representation used to display the similarity or dissimilarity between individual data points in a dataset. Think of it as a map where items that are more alike are placed closer together, and items that are more different are placed further apart.
- LDA (Linear Discriminant Analysis): Separates and classifies samples based on population origin. Linear Discriminant Analysis (LDA) works by finding the lines or directions that best separate different ethnicities based on genetic data. In essence, LDA helps you to draw the best boundaries between different categories based on their features, which makes it easier to classify them correctly. In the real world, this can be used for things like identifying which group a new population belongs to, diagnosing diseases based on patient data, or even recognizing faces in images.
- t-SNE (t-Distributed Stochastic Neighbor Embedding): Another dimensionality reduction technique. t-SNE, or t-Distributed Stochastic Neighbor Embedding, is a machine learning algorithm used to visualize high-dimensional data by reducing it to two or three dimensions, which can be easily plotted and understood. The goal of t-SNE is to take data that has many features, which can be thought of as dimensions in a high-dimensional space, and represent it in a way that preserves the important relationships between data points.
- Dendrogram and Treegram: Shows hierarchical relationships within the populations.
How to Use
- Start with the Montecarlo Configuration to set up your core analysis parameters.
- If desired, configure the Oracle analysis for an alternative perspective.
- In the Analysis Tools Configuration, select which visualization and analysis methods you want to apply to your results.
- Adjust individual settings within each selected method as needed.
- Run your analysis with the chosen configuration.
Remember that each option can significantly impact your results, so it’s important to understand the implications of each setting.